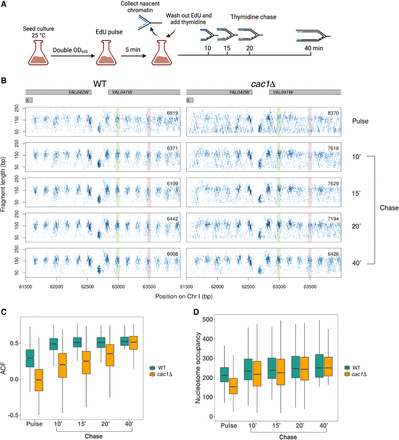
Nascent chromatin occupancy profiling (NCOP) reveals a chromatin maturation defect in cac1Δ cells. (A) Schematic of experimental design for capturing chromatin maturation dynamics behind the replication fork. Panel created with BioRender (https://www.biorender.com). (B) Chromatin occupancy profiles at a representative locus in WT and cac1Δ cells after a 5-min EdU pulse and 10, 15, 20, and 40 min following a thymidine chase. Each dot represents a fragment midpoint, the shading of which is determined by a 2D kernel density estimate. The number of fragments in each window is inset on each panel. Gene bodies are shown in gray on the top. An example of a slow-to-mature nucleosome is highlighted in red, and an example of a fast-maturing nucleosome is highlighted in green. (C) Autocorrelation function (ACF) values at each time point for the 2097 genes with regularly phased nucleosome arrays in mature chromatin, defined as genes with ACF greater than the median in the WT 40-min chase sample. (D) Nucleosome occupancy at all time points for the 41,663 high-confidence nucleosomes, defined as nucleosomes with occupancy greater than the 25th percentile in both WT and cac1Δ cells following the 40-min chase period.











