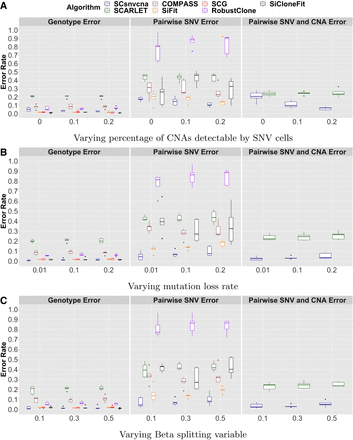
Figure 4.
Group 3: evaluation of methods given biological challenges. Genotype error, pairwise SNV error, and pairwise SNV/CNA error are shown for SCsnvcna, SCARLET, COMPASS, SiFit, SCG, RobustClone, and SiCloneFit on varying percentage of CNAs detectable by SNV cells (A), varying mutation loss rate (B), and varying beta splitting variable (C).











