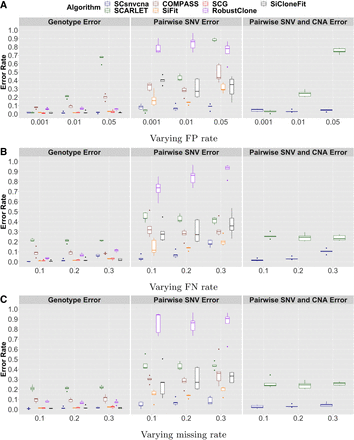
Figure 2.
Group 1: evaluation of methods given varying sequencing errors. Genotype error, pairwise SNV error, and pairwise SNV/CNA error are shown for SCsnvcna, SCARLET, COMPASS, SiFit, SCG, RobustClone, and SiCloneFit on varying FP rate (A), varying FN rate (B), and varying missing rate (C).











