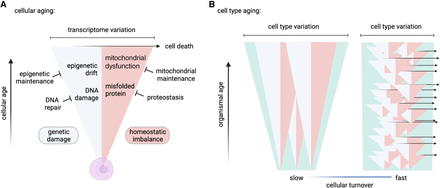
Models of cellular aging and cell type–specific transcriptome variation. Here we present a model depicting the accumulation of transcriptome variation with cellular age. (A) As a cell ages, it accumulates genetic damage (light blue) and cellular homeostasis becomes increasingly imbalanced (pink), either of which can directly or indirectly increase transcriptome variation. The rates of accumulation of damage and loss of homeostasis are counteracted by repair and maintenance processes, which in turn slow the rate of increase in transcriptome variation. As cellular transcriptome variation increases, so does the likelihood of triggering cell death (rightward arrow) (Tower 2015). (B) The model of cellular aging when applied to populations of long-lived cells (slow turnover) and short-lived cells (fast turnover) describes the accumulation of transcriptome variation among a cell type (green). Long-lived cells age together with the organism, and transcriptome variation accumulates among these cells as the organism ages. In short-lived cells, damaged cells are frequently replaced by new cells, which prevents accumulation of variation at the cell type level. Cartoons were created with BioRender (https://www.biorender.com).











