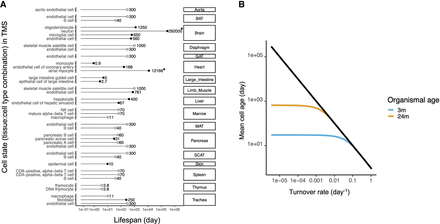
The distribution of cell life span data used in this study and of mean cell age distributions during organismal aging. (A) Summary of cellular life span data used in this study. The cellular life span data in rodents (Sender and Milo 2021) were aligned with cell types in the TMS data (see Supplemental Table S1; Almanzar et al. 2020; Zhang et al. 2021). The left-side texts show cell type names; the right-side texts, tissue names: (BAT) brown adipose tissue, (GAT) gonadal adipose tissue, (MAT) mesenteric adipose tissue, and (SCAT) subcutaneous adipose tissue. Filled circles indicate the 15 cell types with tissue-specific life span data; open circles, without tissue-specific life span estimates that correspond to seven distinct cell types. Note that in two cases, the statistical model used by Sender and Milo (2021) gives life span estimates that exceed the maximum life span of laboratory mice, highlighted with an asterisk for neuron and atrial myocytes. However, these values lead to minimal effects on the mean cell age during organismal aging. (B) Mean cell age as a function of cell turnover rate. The black line denotes the theoretical expectation for a cell with an exponential survival function in an infinitely long-lived organism. More realistically, we assume that cell age follows a truncated exponential distribution, such that the mean cell age is constrained by organismal age. We show the analytical solution of the mean cell age as a function of cell turnover rate for a 3-mo-old mouse (blue) and a 24-mo-old mouse (yellow), using the model in Equation 8 (Supplemental Methods).











