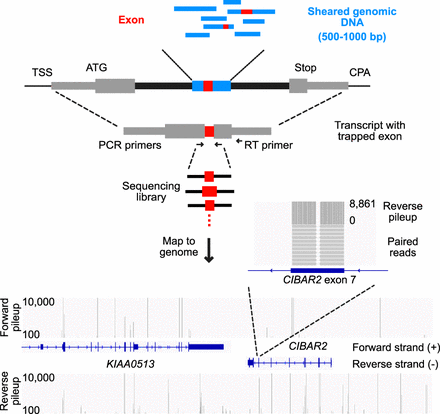
Overview of the genome exon trapping method. Diagram depicting exon trapping approach, sequencing library construction and example sequencing read maps. Sheared genomic DNA library fragments (blue boxes) 500–1000 bp in length are cloned into the middle of the sixth intron from TRA2B (black boxes), in a pcDNA 3.1 vector backbone. First and terminal exons (gray boxes) are labeled with the transcriptional start site (TSS), start codon (ATG), stop codon (Stop), and the cleavage and polyadenylation site (CPA). Internal exons (red boxes) are amplified by RT-PCR, using indicated primers, then sequenced and mapped to the human genome (hg38). Bottom panel shows mapped sequencing read counts (separated into forward and reverse strand pileups) for regions containing KIAA0513 and a portion of CIBAR2 (display region coordinates: Chr 16: 85,062,938–85,134,585). The zoom-in region corresponds to exon 7 of CIBAR2.











