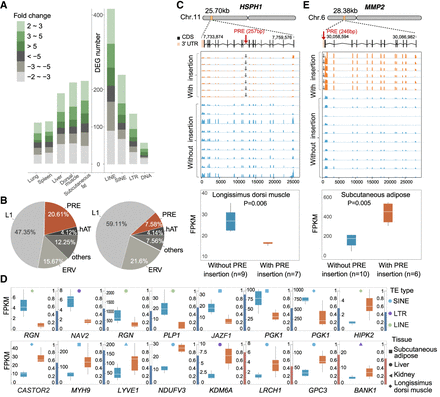
Gene expression differentiation by active TEs. (A, left) The number of differential expression genes affected by TIPs in five tissues (spleen, muscle, subcutaneous adipose, kidney, and liver). (Right) Comparison of the number of differential expression genes affected by different TE types. (B, left) Percentage of major TIP superfamilies or families affecting gene expression. (Right) Percentage of major TE families of pig mobilome. (C) The sketched gene structure of HSPH1, and the comparison of HSPH1 expression levels (FPKM) between individuals with a PRE insertion and individuals without a PRE insertion. (D) The sketched gene structure of MMP2, and the comparison of the MMP2 expression level (FPKM) between individuals with a PRE insertion and individuals without a PRE insertion. (E) Sixteen genes harbor population-specific TE insertions with a presence frequency > 0.4; comparisons of the gene expression level (FPKM) of these genes between individuals with population-specific TE insertions (orange) and without population-specific TE insertions (blue). The red and blue bars on the right side represent the presence frequencies of TE insertions present only in Chinese pigs and European pigs, respectively.











