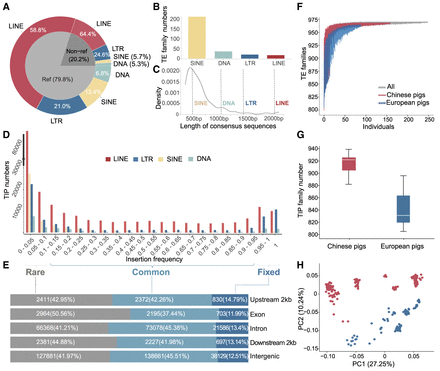
The composition of pig mobilome. (A) Percentage of four major TIP types (DNA transposons, SINE, LINE, and LTR) in the reference (Ref) and nonreference (Non-ref) genome. (B) The number of newly identified TE families in four major TE types. (C) The distribution of the length of newly identified TE families. The vertical dotted lines denote the average length of consensus sequences of four major TE types in Repbase. (D) Frequency distribution of four types of TIPs—LINE, SINE, LTR, and DNA—in the 250 genomes. (E) Proportions of rare, common, and fixed TIPs in intergenic and intragenic regions. TIPs observed in <5% of animals are designated as “rare,” between 5% and 95% as “common,” and >95% as “fixed.” (F) The cumulative number of TE families identified with increasing numbers of individuals by iterative resampling of individuals. Red, blue, and black represent Chinese pigs, European pigs, and all 250 individuals, respectively. (G) Comparison of the number of TE families with TIPs in Chinese and European genomes. Only genomes with sequencing coverage >30× were included (904.01 ± 34.6 in Chinese and 845.6 ± 28.9 in European genomes, respectively). (H) Principal component analysis based on PRE with TIPs. Chinese pigs and European pigs were well separated. Colors represent the populations as indicated in F.











