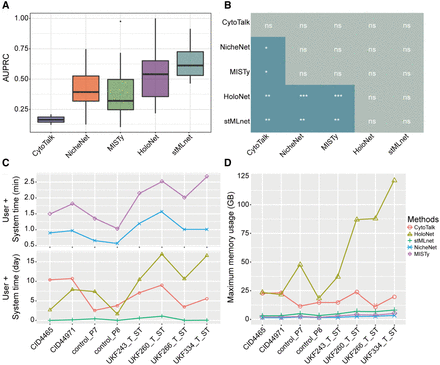
Evaluation and comparison of five L/R-target inference methods using cell line perturbation-expression data sets. (A) Box plot of AUPRC values of each method across all combinations of the input ST data sets and perturbation-expression benchmark data sets. (B) Heatmap of statistical significance of difference of AUPRC values between each pair of two methods. Blue represents that the AUPRC values of the method in row are significantly higher than those of the method in column ([*] P ≤ 0.05, [**] P ≤ 0.01, [***] P ≤ 0.001, [ns] P > 0.05), otherwise for gray. (C) Line chart of running time (user time + system time) of the methods on different data sets. (D) Line chart of maximum memory usage of the methods on different data sets.











