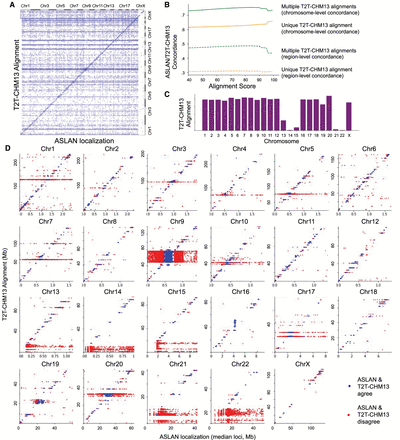
Comparison between ASLAN localizations and T2T-CHM13 alignments. (A) Confusion matrix comparing ASLAN localizations, lifted over to T2T-CHM13 coordinates to T2T-CHM13 alignments of 100-mers extracted from the unmapped reads, binned into 1000 equally sized bins across the genome. (B) Concordance rate between ASLAN localization and T2T-CHM13 alignment versus alignment score, colored by whether or not alignment to T2T-CHM13 was a unique mapping or not. (C) Concordance rate between ASLAN localization and T2T-CHM13 alignment, versus the chromosome to which ASLAN localized. Acrocentric Chromosomes 13–15 and 21–22 show a significantly lower concordance. (D) T2T-CHM13 alignment versus center point of ASLAN localization region, separated by chromosome and colored by whether or not T2T-CHM13 alignment and ASLAN localization were in concordance.











