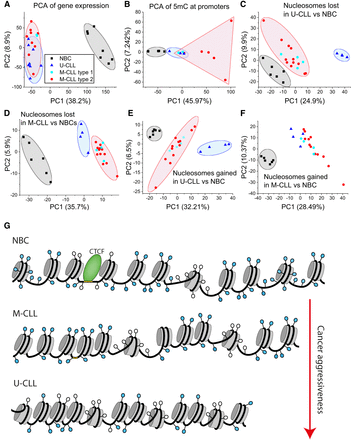
Nucleosome positioning as a new marker to stratify CLL patients. Panels A to F show principal component analysis (PCA) based on gene expression, DNA methylation, and nucleosome occupancy. Each dot represents one sample from NBC (black squares), M-CLL (red circles), and U-CLL (blue triangles). (A) PCA based on gene expression. (B) PCA based on 5mC at promoters. (C) PCA based on nucleosome occupancy at regions with reduced nucleosome occupancy in U-CLL versus NBC. (D) Same as in panel C but for regions with reduced nucleosome occupancy in M-CLL versus NBC. (E) Same as panel C but for regions with increased nucleosome occupancy. (F) Same as panel D but for regions with increased nucleosome occupancy. (G) A scheme of molecular mechanisms of nucleosome repositioning in CLL. CLL B cells have shorter NRL and more partially unwrapped nucleosomes than NBCs. These features are linked to differential DNA methylation, rearrangement of CTCF and other TFs, and active chromatin remodeling. The aberrant nucleosome positioning in CLL is more pronounced for the more aggressive U-CLL subtype compared to M-CLL.











