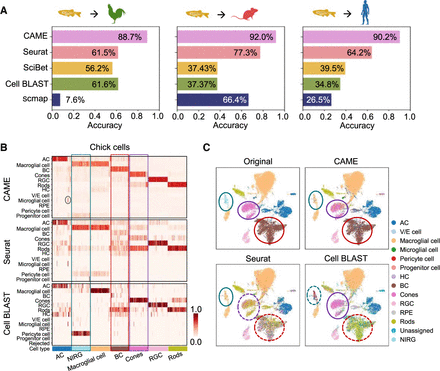
Application of CAME to retina scRNA-seq data between distant species. (A) Performance comparison of CAME and four other approaches, including Seurat, scmap, SciBet, and Cell BLAST, in terms of cell typing accuracy on three pairs of cross-species scRNA-seq data sets about the retina with zebrafish as the reference and with three species (chick, mouse, and human) as the queries. (B) Heatmap comparison of the assignment probabilistic matrices of CAME and another top two methods (Seurat and Cell BLAST) in A for each query cell (column) about the retina with zebrafish as the reference and with chick as the query. For convenience, only 100 cells were subsampled from each cell type to visualize. Note that Cell BLAST only provides the P-value for each query cell to its k (k = 5 by default) nearest reference cells, so the probabilities were computed by averaging the labels of these reference cells with P-values lower than 0.05. (C) UMAP plots of the chick retinal cells colored by the original assignments and the predicted ones by CAME, Seurat, and Cell BLAST using zebrafish as the reference.











