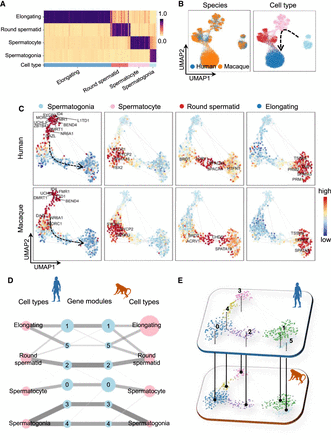
Application of CAME to human and macaque scRNA-seq data during spermatogenesis. (A) The predicted cell-type probabilities for each macaque testicular cell (each column). A maximum of 50 cells was subsampled from each type for visualization. Gene expression in human testis was taken as the reference. Each row indicates a cell type in the human data. (B) The UMAP plots of cell embeddings output by CAME, colored by data sets (left) or cell type (right). (C) 2D visualization of gene embeddings showing the average expression patterns (z-scored across cell types for each gene) of the four stages across spermatogenesis, where each point represents a gene, and the color of each scatter is scaled by the expression level of that cell type in the gene. (D) Abstracted graph of the heterogeneous cell–gene graph. Each node represents a cell type (pink) or a gene module (light blue). The size of a node is scaled by the number of single cells in that type or the number of genes in that gene module. The width of an edge is scaled by either the normalized mean expression levels of a cell type in the connected gene module or the conservancy of inter-species gene modules based on the gene embeddings learned by CAME. (E) Gene modules detected by joint module extraction of genes from humans (above) and macaques (below).











