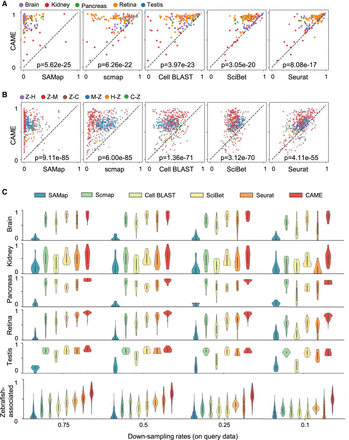
Benchmarking cross-species cell-type assignment performance of CAME. (A,B) Performance comparison of CAME and five other approaches in terms of cell typing accuracy on 139 pairs of cross-species scRNA-seq data sets (A) and on 510 pairs of cross-species scRNA-seq data sets that associated with zebrafish (B), where each point represents a pair of cross-species data sets and is colored by tissue. The notation “X-Y” indicates that X is the reference and Y is the query. (H) Human, (M) mouse, (C) chick, (Z) zebrafish. (C) Performance comparison of the classification accuracies of CAME and the five other methods on different down-sampling rates (0.75, 0.5, 0.25, 0.1) for read counts.











