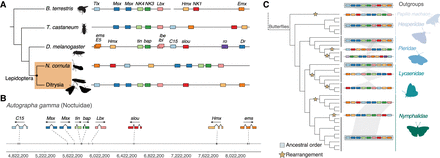
NK gene cluster evolution across Insecta. (A) Comparison of the general structure of the NK gene cluster between representative species for Hymenoptera (B. terrestris), Coleoptera (T. castaneum), Diptera (D. melanogaster), and Lepidoptera. Lepidoptera are shaded in an orange box and split between non-Ditrysia species (N. cornuta) and Ditrysia (represented by 122 species in our data set). (B) Genomic location of NK genes in A. gamma with corresponding exon structures and genomic distances annotated below. Silhouette images of B. terrestris, T. castaneum, and D. melanogaster were taken from PhyloPic (phylopic.org). (C) Left shows the species topology for the 36 butterflies in the data set, along with an outgroup representative. Rearrangements in the NK cluster are annotated on the branches of the tree where they were estimated to have occurred (represented by yellow stars). Black lines spanning tips on the tree group species, which show the same structure and order in the NK gene cluster. The NK gene cluster is represented by colored boxes, in the “canonical” order of Tlx (C15), Msx (Dr), NK4 (tin), NK3 (bap), Lbx (lbe), NK1 (slou), Hmx (NK5), and Emx (ems). Species with the NK genes in this order are shadowed by a blue box. Synteny between the closely linked genes of both Msx (Dr) genes, NK4 (tin), NK3 (bap), and Lbx (lbe) is represented by shaded blocks to show changes in the order and structure of the NK cluster.











