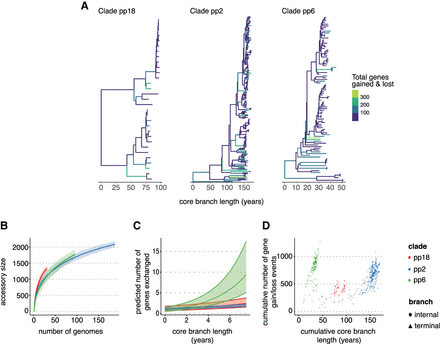
Analysis of the pangenome dynamics of an E. faecalis data set. (A) Phylogenies of three major E. faecalis clades taken from Pöntinen et al. (2021) with branches colored by the number of gene gain and loss events inferred using maximum parsimony. (B) The pangenome accumulation curves of the same clades using the pangenomes inferred using Panaroo in Pöntinen et al. (2021). (C) The predicted slope of the relationship between core genome branch length and the number of gene gain and loss events is inferred by the Panstripe algorithm. (D) The cumulative number of gene gain and loss events versus the cumulative branch length starting from the root node of each tree in A. This is a similar plot to the common “root-to-tip” plot used in phylogenetic dating.











