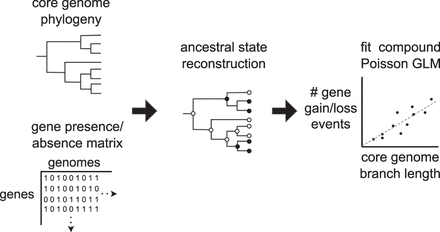
A schematic of the Panstripe algorithm. A binary gene presence/absence matrix and a core genome phylogeny are taken as input. The ancestral states of each gene are then determined before fitting a compound Poisson GLM to compare the core branch length with the number of gene gain and loss events on each branch. Additional terms are included to account for the depth of a branch and whether or not it occurs at the tips of the phylogeny.











