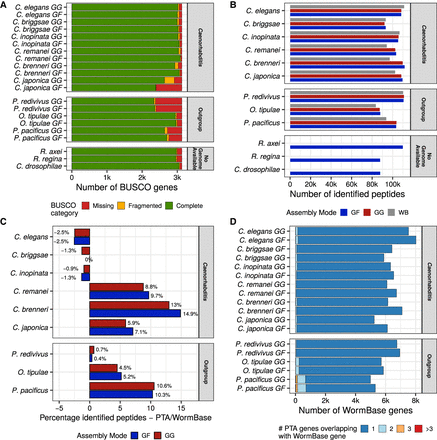
Proteotranscriptomics assembly (PTA) of 12 nematode species. (A) BUSCO metrics for all species and the two assembly modes, GF and GG. (B) Bar plot of mass spectrometry–identified peptides belonging to protein entries of the respective annotation source (GF, GG, and WB) for each species. (C) Peptide identification improvement of GF and GG annotations compared with WormBase annotations. (D) Number of GF and GG proteins with peptide evidence overlapping with the respective WormBase protein annotation for each species. Light blue, orange, and red groups represent WormBase entries that are covered by more than one proteotranscriptomics-validated protein.











