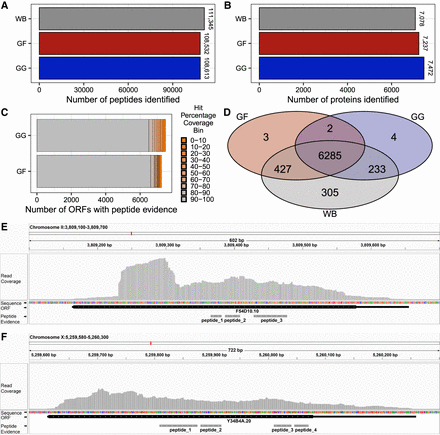
Proteotranscriptomics in C. elegans. (A) Number of peptides identified in the C. elegans WormBase, GF, and GG assemblies. (B) Number of individual proteins with peptide evidence for WormBase, GF, and GG proteomes. (C) Stacked bar plot of all ORFs of the GF and GG assembly with peptide evidence grouped by the level of coverage with the respective WormBase entry. (D) Venn diagram depicting the overlap between the identified proteins using WormBase, GF, or GG assembly as search space for peptide identification. (E) Visualization of the new C. elegans gene F54D10.10 via the Integrative Genomics Viewer (IGV) (Thorvaldsdóttir et al. 2013) aligned to the C. elegans genome sequence. Presented are read coverage (gray peak track), ORF structure (black bar; thick bar represents translated region), and position of peptide evidences (gray bars). (F) Same as E for Y34B4A.20.











