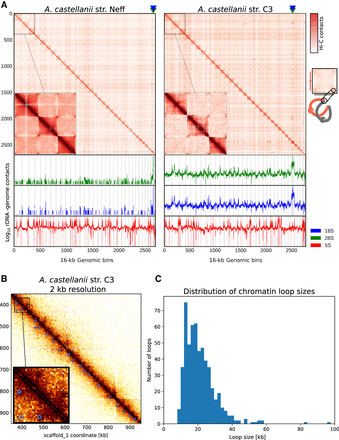
Spatial organization of the A. castellanii genome. (A, top) Whole-genome Hi-C contact maps of the Neff (left) and C3 (right) genomes, with a magnification of the three largest scaffolds. The genomes are divided into 16-kb bins, and each pixel represents the contact intensity between a pair of bins. Each scaffold is visible as a red square on the diagonal. In both strains, there is an enrichment of interscaffold contacts toward telomeres, suggesting a spatial clustering of telomeres, as shown on the model in the right margin. (Bottom) 4C-like representation of spatial contacts between rDNA and the rest of the genome. Scaffolds are delimited by gray vertical lines. Contacts of all rDNAs are enriched toward telomeres. The genomic position of the 18S and 28S genes is highlighted with triangles on the top panel, and the occurrences of 8S rDNA sequences are shown with vertical red lines on the bottom panel. (B) High-resolution contact map for a region of the C3 genome showing chromatin loops detected by Chromosight as blue circles. (C) Size distribution of chromatin loops detected in the C3 strain.











