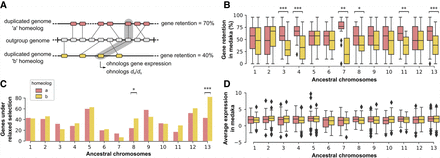
Differences in gene retention, selective pressure, and gene expression between duplicated chromosomes. (A) Schematic example of gene retention calculation. Using an outgroup genome as an approximation of the ancestral gene order, we assess gene retention on each duplicated chromosome in teleost genomes, by 10-gene bins, regardless of their genomic location (Methods). (B) Gene retention on anciently duplicated chromosome copies in medaka, using the spotted gar genome as a proxy for ancestral gene order. Ancestral chromosomes with a significant bias in gene retention on one of the two copies are highlighted ([***] P < 0.001, [**] P < 0.01, [*] P < 0.05, Wilcoxon paired tests with Benjamini–Hochberg correction for multiple testing). (C) Number of genes experiencing relaxed selection compared with their ohnolog across homeologs (Methods; Fisher's exact tests with Benjamini–Hochberg correction for multiple testing, P-values as in B). (D) Average expression across tissues in medaka. No significant differences in expression were detected between genes of duplicated chromosome copies (Wilcoxon paired tests with Benjamini–Hochberg correction for multiple testing, at α = 0.05).











