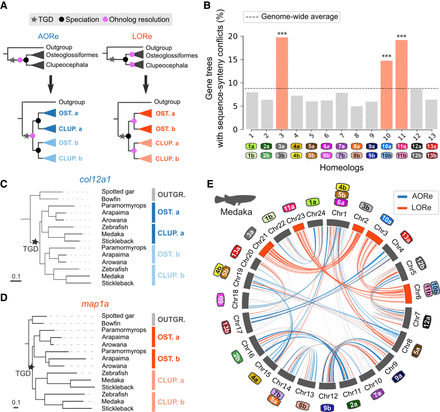
Delayed rediploidization following the TGD. (A) Gene tree topologies expected under the AORe and LORe models. The AORe tree topology assumes that rediploidization was complete before the divergence of Osteoglossiformes and Clupeocephala, initiating “a” and “b” gene sequence divergence before speciation. The LORe tree topology assumes that rediploidization was completed only after the divergence of Osteoglossiformes and Clupeocephala, delaying “a” and “b” duplicated sequence divergence to after speciation. (B) Ancestral Chromosomes 3, 10, and 11 are enriched in sequence-synteny conflicts (Methods, [***] P < 0.001, hypergeometric tests with Benjamini–Hochberg correction for multiple testing). Color labels identify ancestrally duplicated chromosomes as in Figure 1B. (C) Examples of an AORe gene tree. For the col12a1a - col12a1b family, the LORe topology is inconsistent with gene sequence evolution (P = 4 × 10−9, AU-test). (D) Example of a LORe gene tree. For the map1aa - map1ab family, AORe was rejected (P-value = 0.001, AU-test). (E) AORe and LORe gene families visualized on the medaka karyotype. Medaka chromosomes are annotated as numbers, whereas color labels represent ancestral chromosomes (Methods), as in (B). Homeologs 3, 10, and 11 almost entirely rediploidized later than the Osteoglossiformes/Clupeocephala divergence.











