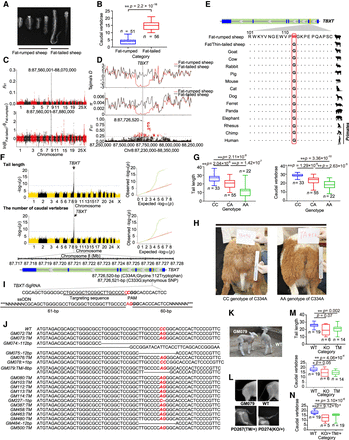
Genome-wide selection sweep test, genome-wide association study (GWAS) analysis, and functional validation of the TBXT gene. (A) Computerized tomography (CT) scanning of the caudal vertebrae number for fat-rumped (Kazakh, Duolang, Bashibai, and Bayinbuluke) and fat-tailed (Hetian and Cele Black) sheep. (B) Boxplots for the number of caudal vertebrae between the fat-rumped and fat-tailed sheep breeds. (C) Genome-wide selection sweep tests (the FST-based and π-ratio-based methods) for the tail configuration between fat-rumped and fat-tailed sheep breeds. (D) Calculation of Tajima's D-values and π and FST values for SNPs in the candidate genomic region Chr 8: 87.56–87.88 Mb between the fat-rumped and fat-tailed sheep breeds. (E) Amino acid (i.e., p. G112W in the TBXT gene) alternation among different mammalian species. (F) Manhattan and Q-Q plots of association signals for the tail length and the number of caudal vertebrae in domestic sheep. The dashed line represents the significance threshold (–log10(0.05/total SNPs) = 8.56). The gene structure of the TBXT gene is shown in green, and the exon regions are shown in blue at the bottom. (G) Boxplots for the tail length and the number of caudal vertebrae associated with the three genotypes of c.334C > A in TBXT in the F2 intercross population of 110 individuals. (H) Different phenotypes in the tail configuration for individuals with the CC and AA genotypes of c.334C > A in TBXT; picture credit: Wen-Rong Li. (I) Sequences of sgRNA targeting exon 2 of the TBXT gene and 123-bp single-strand DNA oligonucleotides (ssODNs) for homologous recombination-mediated repair in CRISPR-Cas9 experimentation. (J) Target sequences in wild-type (WT) and 19 mosaic mutant merino sheep. Target mutations are indicated in the red bold font. (K) Different phenotypes of the tail configuration for WT and mutant merino sheep (GM079); picture credit: Wen-Rong Li. (L) CT scanning of the tail configuration for WT, GM079, and the offspring of GM079. (M) Boxplots for the tail length and the number of caudal vertebrae of the 19 WT sheep, 14 C334A target mutation (TM) merino sheep, and six sheep with a short indel (KO) in the TBXT gene. (N) Boxplots for the number of caudal vertebrae of the 19 WT sheep and 24 offspring of GM079 (i.e., 19 heterozygotes with the C334A target [TM/+] and five heterozygotes with an 8-bp deletion [KO/+]).











