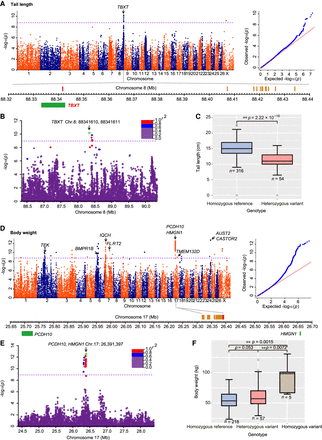
Identification of candidate genes related to the tail length and body weight of domestic sheep. (A,D) Manhattan and quantile–quantile (Q-Q) plots of association signals for the traits of tail length and body weight. The dashed lines colored purple represent the significant thresholds with –log(P-value) = 8.88. SNPs near the peaks with different significant values are marked in red (maximum –log10(P)) and brown (–log10(P) ≥ 7). Functional genes surrounding the peaks are indicated by green boxes. (B,E) Linkage disequilibrium (LD) plots in the regions surrounding the TBXT and HMGN1 genes. (C,F) Boxplots for the tail length associated with SNP (Chr 8: 88,341,610 C/A) and for the body weight associated with SNP (Chr 17: 26,391,397 T/G). The lines in boxes denote the median values; box limits are the upper and lower quartiles, and whiskers show the range of the data; n indicates the number of individuals with the same genotype. Significance of differences between the phenotypic values: (*) P < 0.05, (**) P < 0.01, determined by the Mann–Whitney U test.











