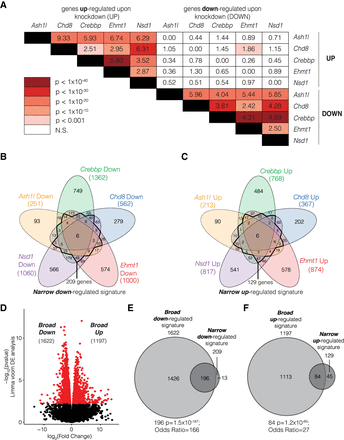
Overlap of genes that are differentially expressed following knockdown of ASD-linked chromatin modifiers. (A) Significance of overlap of down- and up-regulated genes after knockdown. Heatmap indicates significance level by hypergeometric testing. Numbers indicate odds ratio. (B,C) Overlap of genes that are down-regulated (B) or up-regulated (C) by multiple targets. Dark line indicates subset of genes differentially expressed in at least three of five gene sets used to define a “narrow” transcriptional signature. (D) Identification of commonly disrupted genes using limma voom differential expression analysis to generate a “broad” transcriptional signature. (E,F) Overlap of broad and narrow down (E) and up (F) signatures. Overlap significance; hypergeometric tests.











