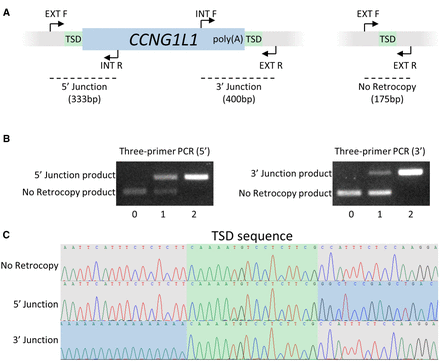
retroCNV validation. (A) Primer design for retroCNV genotyping, with CCNG1L1 as an example. When CCNG1L1 is present, the EXT_F and INT_R primers produce a 333-bp product at the 5′ junction, and the INT_F and EXT_R primers produce a 400-bp product at the 3′ junction. When the retrocopy is absent, the EXT_F and EXT_R primers produce a 175-bp product. (B) Three-primer PCR results for CCNG1L1 at the 5′ and 3′ junctions for individuals with zero, one, and two copies of CCNG1L1. The two external primers EXT_F and EXT_R are included in both reactions, as well as one of the internal primers, INT_F (5′ Junction) or INT_R (3′ Junction). (C) Sanger sequencing results for the CCNG1L1 retroCNV. The TSD is identified as the genomic sequence from the insertion site, which is present at both the 5′ and 3′ ends of the retroCNV.











