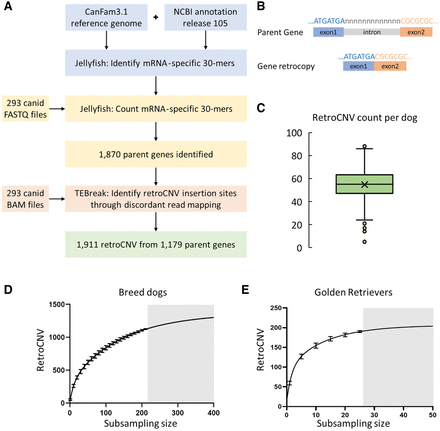
retroCNV discovery. (A) retroCNV discovery pipeline. (B) mRNA-specific sequences are formed owing to intron removal; these sequences are used to identify nonreference retroCNV parent genes in FASTQ reads from genomic DNA sequence. (C) Total nonreference retroCNV count seen in individual domestic dogs (average, 54.1). (D) Number of detected retroCNV insertions with increasing resampling sample sizes in breed dogs. Subsample sizes were selected from one to 210, with a step size of 10, and 100 replicates within each subsample. Bars represent standard deviation of the replicates at each subsample size, and the gray area shows the prediction of increasing the number of dogs beyond the number used in this study. (E) Number of detected retroCNV insertions with increasing resampling sample sizes in Golden Retrievers. Subsample sizes were selected from one to 25, with a step size of five, and 100 replicates within each subsample. Bars represent standard deviation of the replicates at each subsample size.











