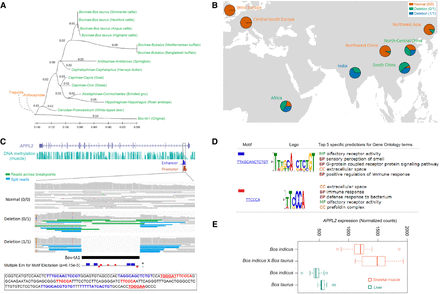
Evolution and function analysis of the Bov-tA1 insertion/deletion in APPL2. (A) The phylogenic tree of Bov-tA1 in APPL2 in species within ruminants. (B) Pie plot of genotype frequencies for Bov-tA1 deletion in APPL2 in cattle according to the geographical distribution. (C) Bov-tA1 location and sequence analyses. (D) GO enrichment analysis of the motifs in Bov-tA1 sequences in APPL2. (E) Boxplot of APPL2 expression levels in skeletal muscle and liver using RNA sequencing data. A number of 15 samples were randomly selected for each population.











