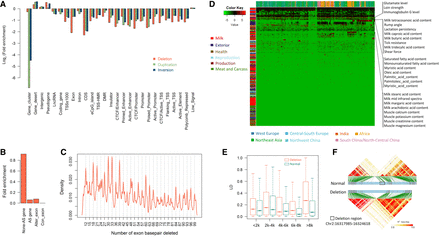
Impacts of the SV on the functional genomic features and QTLs. (A) Enrichment of SVs within various genomic features. (B) Enrichment of SVs within genes with alternative splicing events. (None_AS_gene) Genes with no alternative splicing event, (AS_gene) genes with alternative splicing events, (Alter_exon) alternatively spliced exons, (Con_exon) conserved exons for the genes with alternative splicing events. (C) Density distribution of the exon sequence length deleted. (D) Enrichment of deletions within different QTLs. The log2(fold enrichment) scores of deletions in each QTL for each animal were used to plot this heatmap. (E) SNP LD changes caused by deletions with different lengths. (F) The change of one LD block induced by one deletion.











