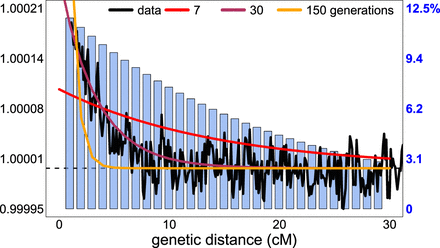
Sampling of DNA segment pairs in fastGLOBETROTTER relative to GLOBETROTTER. Scaled probability (black line), estimating Equation 4 in the Methods, that two segments separated by the given genetic distance are both inferred to share a most recent ancestor with an Irish reference individual, with a key at left. These probabilities are averaged across 20 simulated individuals with European (French) and South Asian (Brahui) admixture occurring 30 generations ago. Blue barplots give the proportion of segment pairs analyzed by fastGLOBETROTTER relative to those analyzed by GLOBETROTTER at each distance bin, with a key at right. To increase computation speed, fastGLOBETROTTER analyzes fewer segment pairs at each distance bin compared with GLOBETROTTER, but it preserves accuracy by analyzing a higher relative proportion of the more informative segment pairs separated by smaller distances. Expected scaled probabilities for three different admixture dates are shown for comparison (legend at top).











