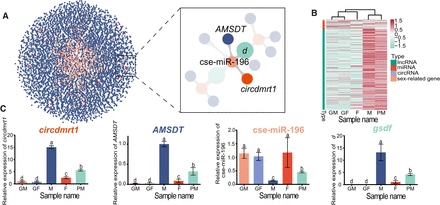
The ceRNAs and sex-related genes with the characteristic expression pattern related to sex determination and differentiation. (A) ceRNA analysis for 48 sex-related genes by Cytoscape. Blue, red, orange, and green nodes represent 4761 lncRNAs, 33 circRNAs, 492 miRNAs, and 48 sex-related genes, respectively. The right image for the ceRNA regulatory network shows enlargement of the boxed region outlined in the left panel. Two related nodes are connected with a gray line. (B) Heat map showing the expression patterns of lncRNAs, circRNAs, miRNAs, and mRNAs in the ceRNETs in the GF, GM, F, M, and PM samples. (C) Expression levels of circdmrt1, AMSDT, cse-miR-196, and gsdf determined by quantitative reverse transcription PCR (qRT-PCR) in the GF, GM, F, M, and PM samples. Actb1 and U6 were used as internal controls. The error bars represent SD. (n = 3.) The different letters abovethe bars denote statistical significance by one-way ANOVA (P < 0.05).











