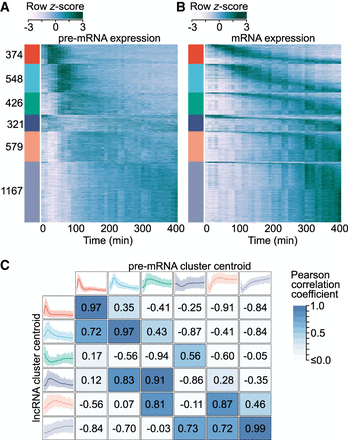
mRNA expression fails to capture gene induction dynamics. (A) Heat map of protein-coding gene induction dynamics. Expression profiles were captured using the first 10 kb of gene introns and z-score normalized. Colored bars (left) indicate cluster membership to one of six clusters obtained through k-means cluster analysis. Clusters are labeled with the number of transcripts in each. (B) Heat map of protein-coding mRNA expression dynamics. Rows within each expression cluster are ranked by the time of peak expression. Rows within A and B correspond to the same genes. (C) Comparison of protein-coding gene pre-mRNA and lncRNA expression dynamics. lncRNA cluster centroids (left) are the same as in Figure 1B, whereas protein-coding pre-mRNA centroids (top) correspond to the colored bars in B. Centroids represent the mean expression of all cluster members, whereas shaded regions represent the 5th–95th percentiles. Pearson correlation coefficients calculated between all lncRNA and protein-coding pre-mRNA centroids are presented.











