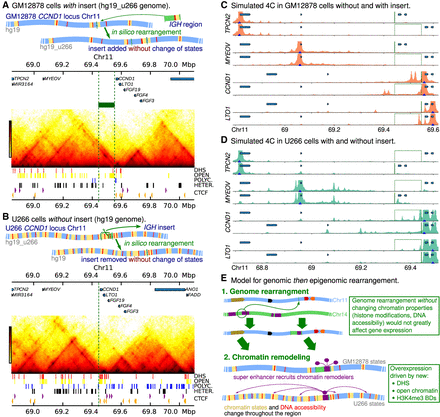
In silico genomic and epigenomic rearrangements. (A, top) Cartoon showing an in silico genome rearrangement where an IGH region is inserted into the GM12878 CCND1 locus, and the GM12878 (healthy) chromatin states are retained (i.e., the hg19_u266 reference genome is used). (Bottom) Hi-C map generated from simulations with this in silico rearrangement. The input data are shown below the map; the green block and dashed lines show the position of the insert. (B, top) Cartoon showing an in silico genome rearrangement where the IGH insert is removed from the U266 CCND1 locus, and the U266 chromatin states are retained (i.e., the hg19 reference genome is used). (Bottom) Simulated Hi-C generated from simulations with this in silico rearrangement. The green dashed line shows where the insert has been removed. (C) Simulated 4C are shown for simulations using GM12878 chromatin states both with and without the insert (position indicated with green lines). Positions of genes are shown by blue blocks (from left to right, TPCN2, MYEOV, CCND1, and LTO1). (D) Simulated 4C are shown for simulations using U266 chromatin states both with and without the insert. (E) Cartoon showing a model where first the genomic rearrangement brings the IGH super-enhancer into the CCND1 TAD, and this then leads to recruitment of chromatin remodelers, etc., to the region and local chromatin states and DNA accessibility are altered. This could be described as an “epigenomic rearrangement.”











