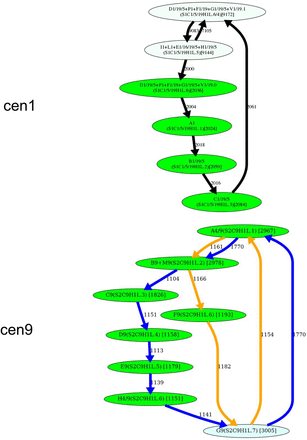
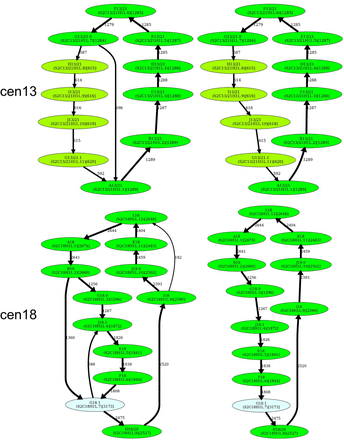
Inferring HORs for cen1, cen9, cen13, and cen18. (First row) Splitting an unbreakable junction monomer in cen1 results in two monomers with an 11-nt difference and transforms the monomer graph of cen1 into a cycle with a single chord. (Second row) The manually inferred HOR of cen9 (McNulty and Sullivan 2018), shown as the blue cycle, is in conflict with the CE postulate because the frequently traversed yellow cycle contains a monomer that does not belong to the blue cycle. (Third row) Splitting an unbreakable junction monomer in cen13 results in two similar monomers with an only 3-nt difference and transforms the monomer graph of cen13 (Fig. 4) into a cycle with a single chord shown on the left. The resulting simplified monomer graph (shown on the right) reveals the canonical 11-monomer HOR in cen13. (Fourth row) Splitting an unbreakable junction monomer in cen18 results in two monomers with only a single-nucleotide difference and transforms the simplified monomer graph of cen18 (Fig. 4) into a cycle with three chords (shown on the left). The resulting simplified monomer graph (shown on the right) reveals the canonical 12-monomer HOR in cen18. (Figure continues on following page.)











