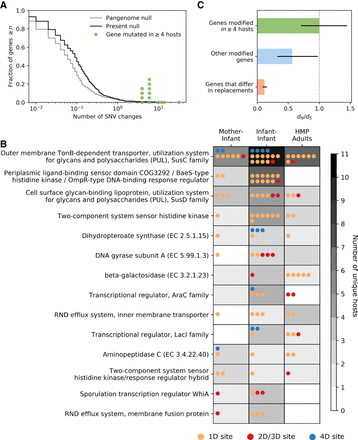
Parallelism of SNV changes across hosts. (A) Mutations are randomized across the metagenomes across hosts to construct two null distributions to assess parallelism (Methods). Observed SNV change counts are shown with green dots for the 14 gene classes found to mutate in parallel in four or more hosts. In the cases in which there are multiple green dots, there are multiple gene classes experiencing the same number of SNV changes. (B) SNV changes in these 14 gene classes found to mutate in parallel in four or more hosts are partitioned by life stage and colored by degeneracy (e.g., 1D indicates a nonsynonymous site, and 4D indicates a synonymous site). Some functional classes are also mutated multiple times within hosts. For inclusivity, the mother–infant age class here includes all QP sample pairs in which the earliest infant time point is taken, irrespective of whether the infant was sampled in the first week of life. This avoids overlapping time points with the infant–infant age group. (C) dN/dS of SNV changes in the 14 gene classes found to mutate in parallel in four or more hosts compared with dN/dS of SNV changes in all other gene classes and dN/dS of sites that differ in strain replacements. The 95% confidence intervals for bootstrapped dN/dS values are reported as black bars.











