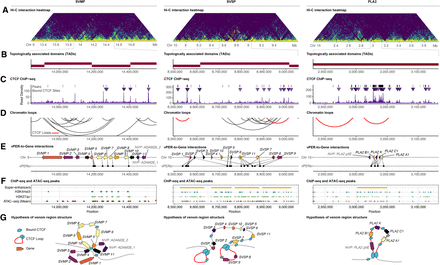
Chromatin structure and organization associated with VG arrays. (A) Hi-C interaction heatmap (10-kb resolution) of 2-Mb regions centered on VG arrays. Brighter colors indicate higher contact frequency. (B) Topologically associated domains (TADs) across VG arrays. (C) CTCF ChIP-seq density, with black bars at the top indicating ChIP-seq peaks and triangles indicating peaks centered on a verified CTCF binding site. (D) Chromatin loops inferred from Hi-C data that span VG arrays. Red loops indicate CTCF–CTCF bound loops, defined by the presence of a CTCF ChIP-seq peak centered around a CTCF motif within 10 kb of both chromatin loop ends. (E) Venom array genes and inferred vPER-promoter interactions. (F) Simplified ChIP-seq and ATAC-seq data, with points indicating the location of ChIP/ATAC-seq peaks and yellow bars indicating SEs if present. (G) Hypotheses for three-dimensional loop structures of VG regions.











