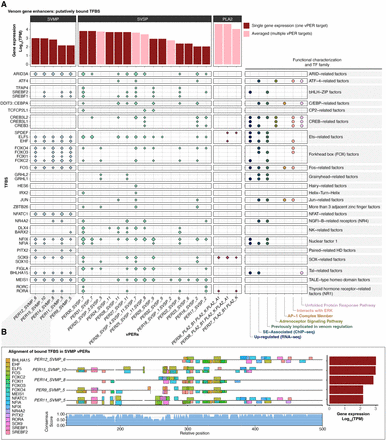
TFBSs in vPERs. TFBSs with evidence for being bound in vPERs of the three major VG families. (A) Diamonds indicate TFBSs inferred to be bound in at least two out of three ATAC-seq replicates based on ATAC-seq footprinting analysis. Size of the diamond corresponds to the number of bound TFBSs inferred in a given promoter (larger = more bound sites). Log-scaled gene expression is shown above, and membership to relevant functional categories and TF families is shown on the right. For vPERs that target multiple genes, the averaged gene expression of target genes is shown (pink bars). Genes for which no bound TFBSs were identified in the promoter are not shown. (B) An example alignment of vPER regions for SVMPs. Colored bars indicate bound TFBSs, with bar height scaled by ATAC-seq footprint score (evidence of site being bound). Bars above and below the line indicate TFBSs inferred on the forward and reverse strand, respectively. Expression of target genes is shown on the right, and the consensus score for the alignment is shown below. SVMP vPERs with no bound TFBSs are not shown.











