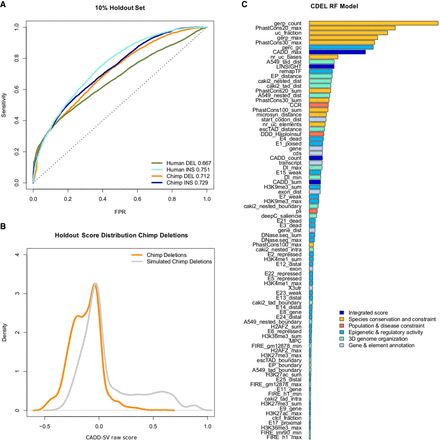
Performance of random forest models trained on proxy-deleterious and proxy-benign SVs. (A) All models show a nonrandom separation of the two classes in a random 10% holdout. Performance is measured as sensitivity over false positive rate (FPR). Note that all training data sets contain a high amount of mislabeled SVs, as a majority of proxy-deleterious SVs is likely to be neutral. (B) Model predictions of the chimpanzee deletion model are shifted toward high-impact SVs in the simulated set of chimpanzee deletions. (C) Representation of feature importance in the chimpanzee deletion random forest model. Note that proxy-pathogenic and proxy-benign sets are length-matched and that length is not used as an explicit feature. Most important contributions come from species conservation (e.g., GERP, phastCons) but also from integrated scores (i.e., CADD or LINSIGHT). Epigenetic features as well as 3D genome architecture features, such as the Directionality Index derived from Hi-C data, also contribute to the most informative features of the models. For a full list of features and explanation of their naming, see Supplemental Table 1.











