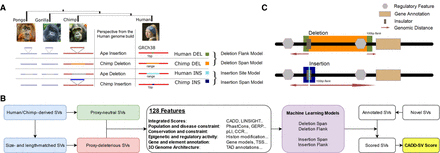
Workflow and training data sets of the CADD-SV framework. (A) Proxy-neutral training data set of CADD-SV. Human- and chimpanzee-derived structural variants (SVs) are considered to be neutral or beneficial if they reached fixation. Therefore, previously identified human- and chimpanzee-derived SVs (Kronenberg et al. 2018) are used as a proxy-neutral training data set. (B) CADD-SV workflow. Size- and length-matched simulated variants are used as a proxy-deleterious training data set. Next, various informative features are annotated and transformed (see Methods; Supplemental Table 1) across span or flank of the variants to train multiple random forest classifiers. Models are used to score user-provided (novel) SVs. For this purpose, variants are annotated, features transformed, and models applied. The maximum value of the flank and span model scores is used as the raw model score. Further, a Phred transformation of the relative rank of the score among gnomAD-SVs provides an easy interpretation of the CADD-SV score. (C) Depiction of implementation of the four models generated from the proxy-neutral and proxy-deleterious variant sets. Whereas deletion of a novel sequence provides information about the deleted sequence in the human genome build, the insertion model relies on the site of integration. Therefore, flanking regions to the SVs are taken into account.











