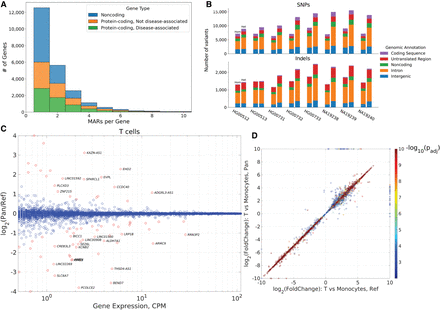
Functional effects of replacing the reference genome with a consensus. (A) Histogram of the number of MARs in the exons of noncoding, protein-coding, and disease-associated genes. (B) Counts of variants in the personal haploid genome that cause mapping errors in the reference, classified by the genomic feature in which the variant is located. For each set of bars, the left bar shows the number of homozygous variants, and the right bar indicates the number of heterozygous variants. (C) The gene expression log2 fold change between the pan-human consensus and the reference genome as a function of the maximum expression in counts per million of the T cell cluster. Red circles indicate genes with an adjusted P-value < 0.1. (D) Comparison of the gene expression log2 fold change between T cells and monocytes in the pan-human consensus and the reference genome. The log2 fold change values were capped between −10 and 10.











