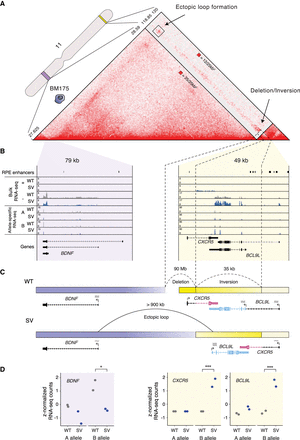
Complex SVs form ectopic loops to simultaneously deregulate BDNF, CXCR5, and BCL9L in cis. (A) Hi-C map highlighting a complex SV on Chromosome 11 in BM175. The SV results in the juxtaposition of the BCL9L/CXCR5 locus (yellow) 90 Mb upstream and the formation of an ectopic loop with BDNF (purple). The genomic regions depicted on the Hi-C maps correspond to the highlighted (purple and yellow) regions on the chromosome ideogram. (B) Gene expression tracks stratified by plus/minus strand (bulk RNA-seq) and A/B allele (allele-specific RNA-seq). SVs affect the B allele only. Dashed vertical lines track the breakpoint positions. (C) Schematic representation of the resolved assembly and the SV-mediated deregulation of BDNF, BCL9L, and CXCR5 in cis, as inferred by Hi-C and RNA-seq data. (D) Allele-specific gene expression of BDNF, CXCR5, and BCL9L. (*) P-value < 0.05, (***) P-value < 0.001.











