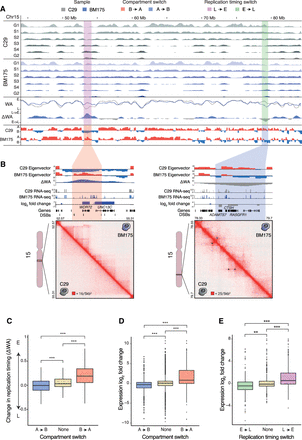
Interplay between compartment switches, replication timing switches, and gene expression in BM175 (daughter clone) compared to C29 (maternal clone). (A) Representative examples of regions with replication timing switches from early → late (green) and late → early (pink) coupled with compartment switches from A → B (blue) and B → A (red), respectively. Percent-normalized density values (PNDVs) for C29 (gray, maternal) and BM175 (blue, daughter) are shown in the top panels. Weighted average values and their difference (ΔWA) pinpoint switches in replication timing, similar to the Hi-C eigenvector tracks below for compartment switches. The switches were evenly distributed genome-wide, irrespective of the presence of DSBs. (B) Alterations in chromatin structure and expression in regions with a combined compartment and replication switch. (Left) B → A and late → early switch accompanied with up-regulation of WDR72 (log2fc = 7.5, P-value = 5 × 10−11) and UNC13C (log2fc = 7.8, P-value = 3 × 10−13), and formation of chromatin loops. (Right) In contrast, an A → B and early → late switch is coupled with down-regulation of ADAMTS7 (log2fc = −3.9, P-value = 1 × 10−15), CTSH (log2fc = −2.2, P-value = 6 × 10−17), and RASGRF1 (log2fc = −2.24, P-value = 0.0002), and loss of chromatin looping. (16/5 kb2) corresponds to 16 normalized counts per 5-kb width. (C) Genome-wide association of compartment switches and changes in replication timing (ΔWA). A → B (n = 298,916) switches have significantly lower ΔWA (E → L) values than regions with no switch (n = 1,912,662), and B → A (n = 317,686) significantly higher (L → E). (***) P-value < 2.2 × 10−16, two-sided t-test. (D,E) A → B (n = 1798) and E → L (n = 327) switches are significantly down-regulating genes compared to regions with no switch, whereas B → A (n = 1063) and L → E (n = 255) up-regulate them significantly. (**) P-value = 3.9 × 10−7, (***) P-value < 7.4 × 10−14, two-sided t-test. Box plots show the median, first, and third quartiles, and outliers are shown if outside the 1.5× interquartile range.











