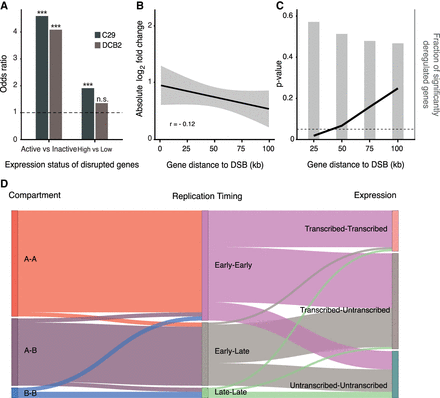
SV-mediated effects on gene expression in cis. (A) DSBs are associated with highly transcribed genes. (Left) Out of 414 DSBs in C29-derived clones, 198 were located within gene bodies; 116 in active, 82 in inactive. The DCB2-derived clone had 137 out of 198 DSBs occurring within gene regions; 80 active and 57 inactive. Genes were defined as active if they had TPM > 1 across all replicates. (Right) Further subdivision of active genes links highly (>50%) transcribed genes with the formation of DSBs. (***) P-value < × 10−5, (n.s.) nonsignificant, Fisher's exact test. (B) Absolute log2 fold change as a function of distance from the closest DSB cluster in BM175 cells. We analyzed in total 274 breakpoints in 182 clustered breakpoint loci. For every DSB cluster, we considered the gene with the highest absolute log2 fold change in a 10-kb sliding window with no overlap. The mean fold change effect at 1 kb versus 100 kb was 0.95 ± 0.35, 95% confidence interval and 0.53 ± 0.33, 95% confidence interval. (C) P-value estimation of the number of significantly deregulated genes as a function of genomic distance from the most proximal TOP2-linked DSB. Background distribution was estimated by randomly shuffling the gene status (deregulated, non-deregulated; see Methods section). Bar plots depict the fraction of deregulated genes within the marked genomic window. (D) Effects of active chromatin, replication timing and gene expression on the formation of TOP2-linked, induced SVs (n = 90). Intermediate replication timing states (n = 214) are excluded for visual simplicity. (See also Supplemental Fig. S6.)











