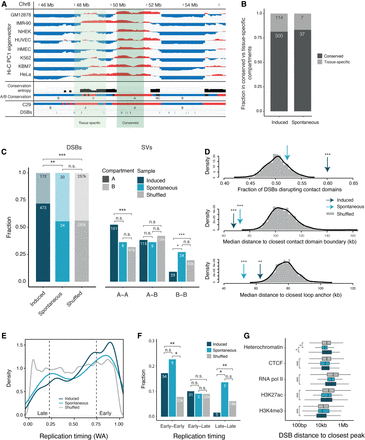
Genome architecture features at SV sites. (A) DSBs associated with conserved and tissue-specific compartments. PC1 eigenvectors from eight human cell lines (Rao et al. 2014), with signs adjusted so that positive values correspond to A (red) and negative values to B (blue) compartments. A/B conservation track annotates every 100-kb bin to a compartment it was found to be in the same state in more than half of the cell lines. Compartment entropy reflects the compartment concordance across the cell lines, with higher values representing less conserved compartments. DSBs in tissue-specific compartments overlap nonconserved regions or regions where the C29 compartment annotation disagrees with the A/B conservation track. (NC) Not conserved. (B) Distribution of DSBs in conserved and tissue-specific compartments between induced and spontaneous rearrangements; 114 out of 614 (18.6%) and seven out of 37 (18.9%) occurred in tissue-specific compartments. (C, left) Induced DSBs (n = 610) are highly enriched within A compartments compared to the spontaneous (n = 44) and shuffled (n = 588,388) set. Fisher's exact test. (Right) SVs are enriched in A to A and depleted in B to B in the induced SV set compared to the shuffled, whereas spontaneous SVs are evenly distributed across A and B compartments. Post-hoc Fisher's exact test, FDR corrected. (D) Observed and expected distribution of DSBs across features that reflect genome organization. (Top) Induced DSBs are enriched inside contact domains (n = 6165), whereas spontaneous are enriched in interdomain loci. Both of them occur significantly closer to insulating factors, such as contact domain boundaries (middle, n = 12,326) and chromatin loop anchors (bottom, n = 17,374) than the shuffled sets. (E) Replication timing weighted average (WA) values at DSBs show enrichment of induced (dark blue, n = 609) and spontaneous (light blue, n = 44) breaks during early replication compared to the permuted set (gray, n = 586,044). (F) SVs occur significantly more frequently between early-to-early replication domains for both induced and spontaneous rearrangements. In contrast, late-to-late replication timing SVs are depleted for induced and enriched for spontaneous SVs. Post-hoc Fisher's exact test. (G) Distance from DSBs to the closest ChIP-seq peak for H3K4me3 (n = 122,258), H3K27ac (n = 103,673), RNA pol II (n = 59,021), and CTCF (n = 43,285) peaks compared to the permuted ones (t-test). Box plots show the median, first, and third quartiles, and outliers are shown if outside the 1.5× interquartile range. (*) P-value < 0.05, (**) P-value < 0.01, (***) P-value < 0.001.











