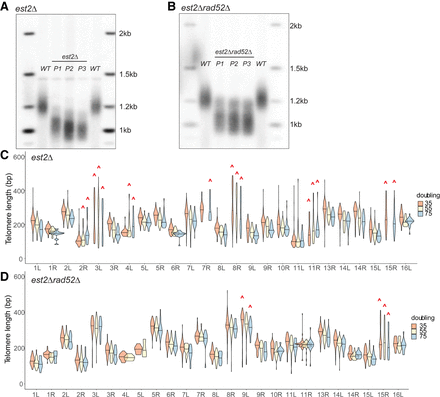
Telomere shortening and outlier elongation events in a telomerase-null, est2Δ background at individual telomeres over time. Freshly generated est2Δ and est2Δ rad52Δ haploids were grown up (P1, 35 doublings) and passaged twice (P2, 55 doublings; P3, 75 doublings) to allow for telomere shortening (see Methods). Telomere length was measured by Southern blot hybridized with the Y′ probe and by nanopore sequencing on the same samples. (A) Southern blot of WT, P1, P2, and P3 est2Δ hybridized with the Y′ probe. (B) Southern blot of WT, P1, P2, and P3 est2Δ rad52Δ hybridized with the Y′ probe. (C,D) Individual telomere length of est2Δ (C) and est2Δ rad52Δ (D) cells determined by nanopore sequencing of P1, P2, and P3. We calculated the sample standard deviations of (σ) of each telomere at every passage and in the wild-type parental strain. Telomeres for which the standard deviation was greater than that of the wild type are considered outliers and are marked with a red caret. At P2, 55 doublings, in the est2Δ sample, not enough data were collected at TEL07R and TEL15R to plot, and at P3, 75 doublings, in the est2Δ rad52Δ sample, not enough data were collected at TEL04L and TEL05L to plot.











