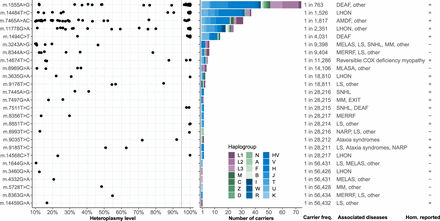
Known pathogenic variants in gnomAD. Shown are the 26 pathogenic variants observed in gnomAD along with their heteroplasmy levels, haplogroup distribution, carrier frequency, MITOMAP-curated disease phenotypes, and indicator showing whether disease occurs at homoplasmy (Hom. reported; note this includes variants only associated with disease at homoplasmy, or at both homoplasmy and heteroplasmy). The carrier frequency is calculated as the high-quality allele count divided by the number of individuals with high-quality sequence at the position. The dark gray line at the 95% heteroplasmy level represents the threshold used to define homoplasmic variant calls. Haplogroups are ordered by their position in the phylogenetic tree and colored by their association with African (purple), Asian (green), or European (blue) ancestry. (AMDF) Ataxia, myoclonus, and deafness, (COX) cytochrome c oxidase, (DEAF) maternally inherited deafness or aminoglycoside-induced deafness, (EXIT) exercise intolerance, (LHON) Leber Hereditary Optic Neuropathy, (LS) Leigh syndrome, (MELAS) mitochondrial encephalomyopathy, lactic acidosis, and stroke-like episodes, (MERRF) myoclonic epilepsy and ragged red muscle fibers, (MLASA) mitochondrial myopathy, lactic acidosis, and sideroblastic anemia, (MM) mitochondrial myopathy, (NARP) neurogenic muscle weakness, ataxia, and retinitis pigmentosa, (SNHL) sensorineural hearing loss, (other) other phenotypes listed for this variant in MITOMAP.











