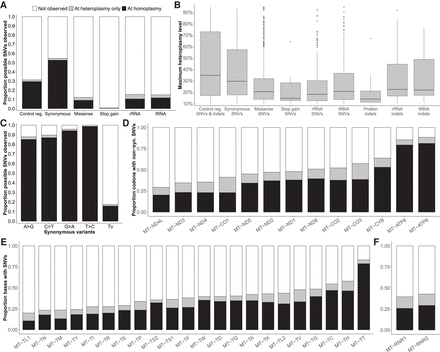
Patterns of variation in the mtDNA in gnomAD. (A) The bar chart shows the proportion of possible SNVs observed, partitioned into those observed at homoplasmy (black), only at 10%–95% heteroplasmy (gray), or not observed (white). (B) The box plot shows the maximum heteroplasmy of variants observed only at heteroplasmy. Protein indels include frameshift and in-frame variants. “Control reg.” represents the noncoding control region m.16024-576 in A and B. (C) The bar chart shows the proportion of possible synonymous variants observed in gnomAD for transversions (Tv) and all possible transitions (A > G, C > T, G > A, T > C) on the reference strand. (D) The bar chart shows the proportion of codons in protein-coding genes with nonsynonymous SNVs observed. (E,F) The proportion of bases in tRNA and rRNA genes with SNVs. Panels C–F follow the color legend in A.











