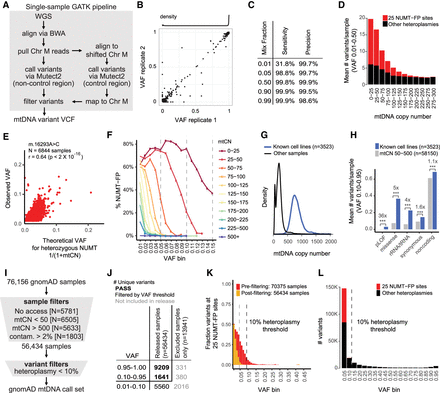
mtDNA call set is designed to exclude NUMT-derived false positives (NUMT-FPs), cell line artifacts, and contaminants. (A) Schematic shows GATK pipeline for calling mtDNA variants in single WGS samples. The control region spans the artificial break in Chromosome M sequence. (B) Reproducibility of GATK pipeline on 91 WGS replicate samples shows 99.3% concordance of calls (2533/2551), and density plot at top shows 87% variants are homoplasmic. (C) Accuracy of single-sample pipeline in samples with mtCN > 500 based on “in silico” mixing data. Note these are valid only for samples with high mtCN. (D) Bar chart shows that the mean number of putative heteroplasmies per sample depends on mtDNA copy number (mtCN), as does the subset occurring at 25 validated NUMT-FP sites (red). (E) Scatterplot shows the observed VAF for a single NUMT-FP (m.16293A > C) across 6844 samples versus the theoretical VAF if the NUMTs were heterozygous and all reads misaligned to the mtDNA. (F) Plot shows VAF levels for NUMT-FP sites decrease with mtCN (colored lines). Y-axis indicates the percent of detected variants that occur at 25 NUMT-FP sites. (G) Density plot shows mtCN for known cell lines and all other samples. (H) Bar plot shows that known cell lines have increased number of heteroplasmic variants in all categories compared to samples with mtCN 50–500 (enrichment shown with *** indicates P-value < 1 × 10−5 based on Fisher's exact test); pLOF indicates predicted loss-of-function. (I) Schematic shows steps for combining and filtering single-sample variant calls into the gnomAD mtDNA call set, designed to exclude NUMT-derived false positives, cell line artifacts, and contaminants. (J) Number of unique variants that pass filters (bold black) versus those filtered out based on VAF (black) or not released (gray). The 19,137 variants are partitioned into mutually exclusive categories; for example, VAF 0.10–0.95 excludes variants also detected VAF 0.95–1.00. (K) For each VAF level, bar chart shows the fraction of variants at 25 NUMT-FP sites before sample filtering (red) or after filtering (orange, shown overlaid). (L) Histogram of VAF (after sample filtering) shows that below 10% VAF, there are a large number of variants and a substantial fraction present at 25 validated NUMT-FP sites (red). X-axis label indicates upper bound of VAF bin.











