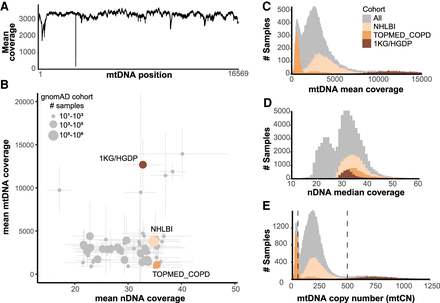
Coverage statistics for 70,375 gnomAD WGS samples. (A) Per-base mean depth of coverage across mtDNA, with coverage dips at positions 303–315 (Tan et al. 2016) and 3107 (Bandelt et al. 2014) due to homopolymeric tract and Chr M reference deletion, respectively. (B) For each cohort within gnomAD, a scatterplot shows the mean nuclear (nDNA) and mtDNA coverage ± standard deviation. Three example cohorts are shown in color: 1000 Genomes and Human Genome Diversity Project cell lines (1KG/HGDP), NHLBI, and TOPMed Chronic Obstructive Pulmonary Disease (TOPMED COPD). (C) Histogram shows mean mtDNA coverage for all samples, and overlaid histograms show three selected cohorts (806 outliers with coverage 15,000–97,000 excluded). We note mean and median mtDNA coverage statistics are extremely similar (Pearson's r = 0.99997). (D) Histogram shows median nDNA coverage for all samples, and overlaid histograms show three selected cohorts (84 outliers with coverage 60–94 excluded). (E) Histogram shows mtDNA copy number per cell (2 × mean mtDNA coverage/ median nDNA coverage) for all samples, and overlaid histograms show three selected cohorts (223 outliers with mtCN 1250–7000 excluded). Only samples with mtCN 50–500 (dashed lines) were included in the released mtDNA call set (56,434/70,375).











