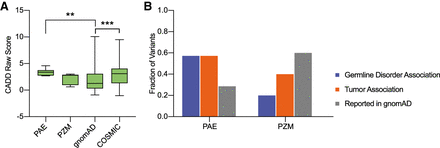
Predictive deleteriousness of variants and association with germline disorders and tumors. (A) Four groups of exonic single-nucleotide substitutions were analyzed: mutations selected from all sequenced libraries as a potential paternal age effect (PAE; n = 7), mutations selected from all sequenced libraries as a potential postzygotic mosaic (PZM; n = 5), exonic single-nucleotide substitutions from the gnomAD v2.1.1 database (gnomAD; n = 876), and exonic single-nucleotide substitutions from the COSMIC v90 database (COSMIC; n = 397). Boxplot comparison of the CADD raw scores obtained for each variant group. A higher score reflects a higher probability of a deleterious effect. Pairwise testing was performed using a Mann–Whitney U test (multiple comparison correction was applied), and only significant differences are marked: (**) P-value < 0.01, (***) P-value < 0.001. (B) Fraction of variants associated with germline disorders according to ClinVar; fraction of variants associated with tumors according to the COSMIC database; and fraction of variants reported in the gnomAD database.











